進化発生ゲノミクス研究グループ
研究紹介
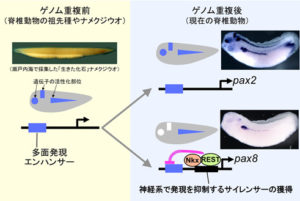
私達の研究室は、「生きた化石」と呼ばれるナメクジウオや様々な生態を持つカエルを材料に、発生制御遺伝子の比較機能解析を通じて、脊索動物のゲノム進化と環境適応の分子メカニズムを研究しています。また、トランスジェネシス技術やゲノム編集技術を用いて、有用な遺伝子組換えツメガエルを作出し、それらを用いて器官(眼や脳、脊髄等)の再生や遺伝性疾患の発症に関わるゲノム・エピゲノム制御の研究をおこなっています。学部生に限らず、他大学からの大学院生、ポスドクなどのメンバーも募集中です。

最近のトピック
スタッフ
*「広島大学研究者総覧」では教員の連絡先、教育担当状況、研究業績などが閲覧できます(公開項目は教員によって異なります)。

博士研究員 林 舜 博士(理学)

技術員 鈴木 菜花
学生・大学院生
研究指導学生:研究課題
Bagus Priambodo(D3):「温泉ガエル」リュウキュウカジカガエルにおけるゲノム多様性と淘汰に関する研究
吉田 真菜(D2):エピジェネティック制御を介した炎症シグナルによる再生プログラムの活性化機構
浅枝 優花(D1):リュウキュウカジカガエルの環境適応に関わる遺伝的変異の探索
荻野 ひなよ(M2):リュウキュウカジカガエル・カジカガエルの初代培養細胞の確立
國重 成恵(M2):アフリカツメガエルにおける疾患関連変異のゲノムワイドな同定
木根森 一仁 (M1):脊椎動物の双眼形成における眼形成遺伝子raxの発現制御メカニズムの解明
島本 百香(M1):ネッタイツメガエル近交系における表現型多型の遺伝的基盤の解明
原田 一平(M1):日本産無尾両生類における幼生の温度嗜好性と行動様式の比較研究
廣田 雅哉(M1):
井上 誓(B4):
鈴木 崇生(B4):
黒川 晃宏(B4):
中崎 真利(B4):
グループからのメッセージ
私たちのラボでは、次世代シーケンサー解析や比較ゲノム解析、高効率トランスジェネシス、ゲノム編集、蛍光タンパク質を用いた生体イメージング、クロマチン免疫沈降等、ウェットからドライまで様々な先鋭的方法を駆使しています。学部生に限らず、他大学からの大学院生、ポスドクなどのメンバーも募集中です。先端的な技術を使ったゲノム進化研究に取り組んでみたい方、野生両生類種の生態から分子まで追究したい方、ヒト先天異常に関する基礎研究に取り組みたい方等、興味があればぜひご連絡ください。見学はいつでも歓迎します。



